- Name
- Jill Rosen
- jrosen@jhu.edu
- Office phone
- 443-997-9906
- Cell phone
- 443-547-8805
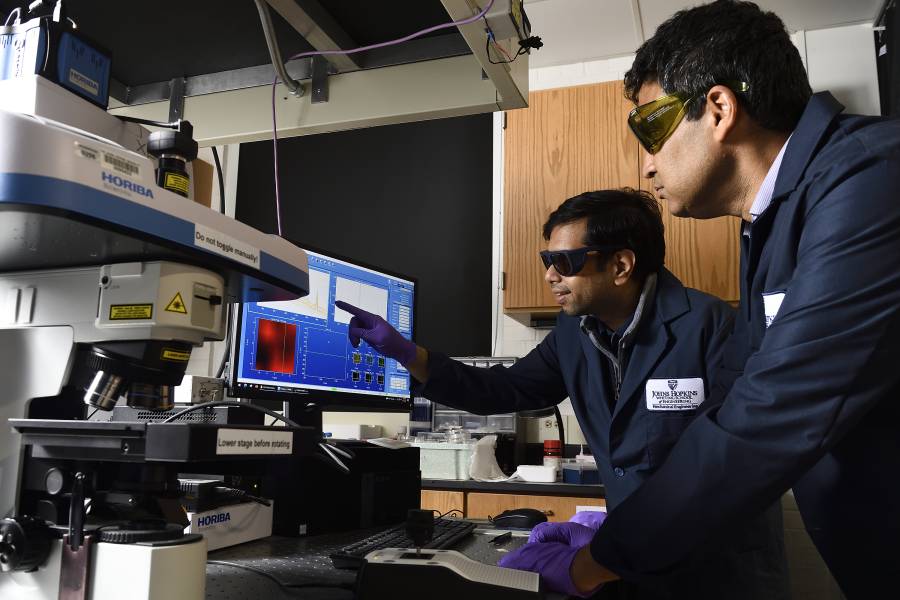
A COVID-19 sensor developed at Johns Hopkins University could revolutionize virus testing by adding accuracy and speed to a process that frustrated many during the pandemic.
In a new study published today in Nano Letters, the researchers describe the sensor, which requires no sample preparation and minimal operator expertise, offering a strong advantage over existing testing methods, especially for population-wide testing.
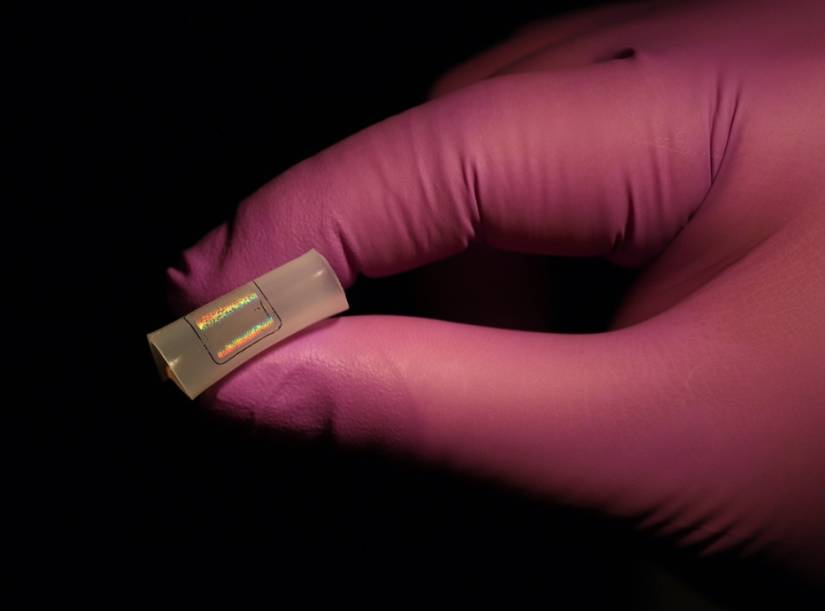

Image credit: Kam Sang Kwok and Aishwarya Pantula /Johns Hopkins University
"The technique is as simple as putting a drop of saliva on our device and getting a negative or a positive result," said Ishan Barman, an associate professor of mechanical engineering, who along with David Gracias, a professor of chemical and biomolecular engineering, are senior authors of the study. "The key novelty is that this is a label-free technique, which means no additional chemical modifications like molecular labeling or antibody functionalization are required. This means the sensor could eventually be used in wearable devices."
Barman says the new technology, which is not yet available on the market, addresses the limitations of the two most widely used types of COVID-19 tests: PCR and rapid tests.
PCR tests are highly accurate, but require complicated sample preparation, with results taking hours or even days to process in a laboratory. On the other hand, rapid tests, which look for the existence of antigens, are less successful at detecting early infections and asymptomatic cases and can lead to erroneous results.
The sensor is nearly as sensitive as a PCR test and as convenient as a rapid antigen test. During initial testing, the sensor demonstrated 92% accuracy at detecting SARS-COV-2 in saliva samples—comparable to that of PCR tests. The sensor was also highly successful at rapidly determining the presence of other viruses, including H1N1 and Zika.
The sensor is based on large area nanoimprint lithography, surface enhanced Raman spectroscopy (SERS), and machine learning. It can be used for mass testing in disposable chip formats or on rigid or flexible surfaces.
Key to the method is the large-area, flexible field enhancing metal insulator antenna (FEMIA) array developed by the Gracias lab. The saliva sample is placed on the material and analyzed using surface-enhanced Raman spectroscopy, which employs laser light to examine how molecules of the examined specimen vibrate. Because the nanostructured FEMIA strengthens the virus's Raman signal significantly, the system can rapidly detect the presence of a virus, even if only small traces exist in the sample. Another major innovation of the system is the use of advanced machine learning algorithms to detect very subtle signatures in the spectroscopic data that allow researchers to pinpoint the presence and concentration of the virus.
"Label-free optical detection, combined with machine learning, allows us to have a single platform that can test for a wide range of viruses with enhanced sensitivity and selectivity, with a very fast turnaround," said lead author Debadrita Paria, who worked on the research as a post-doctoral fellow of Mechanical Engineering.
The sensor material can be placed on any type of surface, from doorknobs and building entrances to masks and textiles.
"Using state of the art nanoimprint fabrication and transfer printing we have realized highly precise, tunable, and scalable nanomanufacturing of both rigid and flexible COVID sensor substrates, which is important for future implementation not just on chip-based biosensors but also wearables," said Gracias.
He says the sensor could potentially be integrated with a hand-held testing device for fast screenings at crowded places like airports or stadiums.
"Our platform goes beyond the current COVID-19 pandemic," said Barman. "We can use this for broad testing against different viruses, for instance, to differentiate between SARS-CoV-2 and H1N1, and even variants. This is a major issue that can't be readily addressed by current rapid tests."
The team continues working to further develop and test the technology with patient samples. Johns Hopkins Technology Ventures has applied for patents on the intellectual property associated with it and the team is pursuing license and commercialization opportunities.
Authors include Kam Sang (Mark) Kwok, a graduate student in Chemical and Biomolecular Engineering; Piyush Raj, a graduate student; and Peng Zheng, a post-doctoral fellow in Mechanical Engineering.
The research was supported by a National Science Foundation's Early-concept Grants for Exploratory Research (EAGER) program and the National Institute of Health Director's New Innovator award.