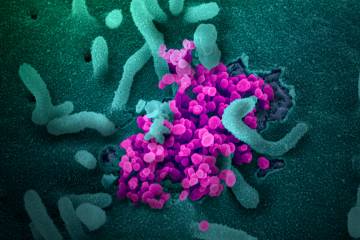
Viruses evolve over time, undergoing genetic changes, or mutations, in their quest to survive. Some viruses produce many variations, others only a few. Fortunately, SARS-CoV-2, the novel coronavirus that causes COVID-19, is among the latter. This is good news for scientists trying to create an effective vaccine against it.
"The virus has had very few genetic changes since it emerged in late 2019," says Peter Thielen, a molecular biologist at the Johns Hopkins Applied Physics Laboratory and JHU Doctor of Engineering candidate, who, with colleagues from other areas of the Hopkins research community, has been sequencing the viral genome to better understand its makeup. "Designing vaccines and therapeutics for a single strain is much more straightforward than a virus that is changing quickly."
SARS-CoV-2 first appeared in China in December before rapidly spreading around the world. In a scant few months, it has sickened more than 7.25 million people worldwide, killing more than 410,000 and counting. In many affected countries, including the United States, the pandemic has prompted extreme mitigation measures to contain it, including lockdowns, extensive quarantines, and social distancing and mask-wearing—restrictions many experts believe will not entirely disappear anytime soon.
"It isn't going to be possible for us to truly be able to return to normal until we have a vaccine," says Winston Timp, assistant professor of biomedical engineering in the Whiting School of Engineering, and who, along with Professor of Medicine Stuart Ray, is leading the Hopkins viral genomics effort. "The low mutation rate of the virus means it should be possible to generate a successful vaccine," he says, adding it also could boost efforts to develop potential treatments for the disease.
Coronaviruses—of which there are hundreds, most of them occurring in animals—typically mutate more slowly than many other viruses. Influenza, for example, mutates quickly, which is why people must be inoculated annually against changing flu strains.
Data from SARS-CoV-2 samples the researchers have examined from the Baltimore and Washington region are similar to those from other parts of the world. "So far, the genetic changes accumulating as the virus spreads are not resulting in different strains of the virus," Thielen says.
This is important because a successful vaccine strategy must account for mutations in order to provide broad protection. "Influenza has a lot of very unique ways of changing over a short period of time, and it does so on local and global scales every flu season," Thielen says. "SARS-CoV-2 is almost the opposite so far—it is changing slowly, and because there is no existing immunity to the virus, it doesn't have any evolutionary pressure to change as it spreads through the population."
Timp agrees. "It's hard to make the right vaccine for flu because there are different strains that circulate every year," he says. "With SARS-CoV-2, there are some small mutations, but nothing to lead us to suspect that if you have immunity here in Maryland that you won't have it anywhere else."
Scientists are focusing on the "spike" protein of the virus, the part that docks with human cells and allows entry. "A vaccine that blocks the virus's ability to infect a cell would be highly effective, since there would be no ability for the virus to generate an active infection to cause or spread disease," Thielen says. "There are very small regions of the spike protein that make direct contact with the receptor on a human cell, and these are the highest likelihood targets for vaccine developers. To date, no changes have been observed to these parts of the virus in any of the more than 20,000 samples that have been sequenced globally."
When a virus replicates—makes copies of its genome within a cell—it uses an enzyme called a polymerase. Some viruses have very accurate versions of these, while others, such as HIV or influenza, do not, Thielen says. "Coronavirus polymerases have something we call 'proofreading activity,' which is exactly like it sounds," he says. "Once a genome is replicated, the enzyme will identify mistakes it has made and correct them. Sometimes it still gets things wrong, however, and small changes can occur."
JHU scientists say they have seen fewer than two dozen mutations between the current versions they are studying and original viral isolates from China, which is a very small number. "This means a vaccine will probably work against all of them," Timp says.
It's still unclear, however, how long immunity will last against this virus, regardless of whether it arises from having recovered from illness or through vaccination.
"The first step is to understand the immune responses required to clear the virus, or protect against it," says Heba Mostafa, an assistant professor of pathology in the School of Medicine who is providing viral material to APL researchers as well as sequencing viral samples in her own lab. She also developed a coronavirus screening test with JHU microbiologist Karen Carroll. "With a stable virus, reinfection will be less likely, or could be less aggressive. This is all helpful, especially in designing a vaccine."
It will be important to monitor mutations that accumulate over time, especially as immunity increases in the population, Thielen says. To that end, APL scientists are working as part of a large group across JHU to characterize the virus's genomic diversity and share information through a multidisciplinary effort, the COVID-19 Research Response Program.
In addition to APL, the initiative includes participants from Johns Hopkins Hospital, the Bloomberg School of Public Health, the Whiting School of Engineering's Department of Computer Science and Department of Biomedical Engineering, which is shared by the engineering school and the School of Medicine. "Data generated from our sequencing efforts gets uploaded to global data repositories, so that all researchers can study the virus at the same time," Thielen says.
The researchers are aware that most people won't feel safe until there is a vaccine. Thielen has friends who have become sick, or who have lost loved ones. And he has a family at home—including two young children—to protect. Nevertheless, based on evidence about the virus generated thus far, he believes a vaccine will be forthcoming.
"As time goes on, it is likely that all of us will have a personal connection to someone who has been impacted by the virus," he says. "What we observe in the local and global data provides excellent reassurance that we are on the right track."
Posted in Health, Science+Technology