Johns Hopkins biomedical engineer Natalia Trayanova builds computer models of real, troubled hearts. The pulse of her project beats from a cluster of white buildings packed with electronic processors and storage on Johns Hopkins' Bayview campus.
Backed with this computational power, Trayanova and her team pursue treatments for heart disease that are more effective because they're tailored to the specific patient and tested on that patient's virtual heart.
Trayanova is one of hundreds of Johns Hopkins scientists in a range of fields whose work is supported by the Maryland Advanced Research Computing Center, a joint effort of Johns Hopkins and the University of Maryland, College Park backed by $30 million in state funding. MARCC has been operating since the summer of 2015 and is helping to produce results for scientists working at computer screens miles away, tapping into its 22,000 processors and 17 million gigabytes of storage.
The center, the 17th largest academic computing complex in the country, has allowed research groups to save the money and space they would otherwise use to build their own computing centers. Investigators on projects as diverse as treating heart disease, improving wind power, and modeling the molecular structures of proteins say MARCC makes a big difference.
"In a very short time, MARCC has become one of the most utilized research facilities at Johns Hopkins" and the University of Maryland, says Jaime Combariza, a Hopkins research professor in Johns Hopkins' Department of Chemistry and director of MARCC. "I think it's fair to say that MARCC has exceeded just about everyone's expectations."
Trayanova, a professor of biomedical engineering, said that for her project, MARCC is "absolutely essential. This is capital letter, bold and underlined. ... MARCC to us is like a godsend."
Trayanova's group recently completed a clinical trial of 10 patients who were suffering persistent heart arrhythmias and completed treatments that were tested using the virtual heart models. She's still waiting to see results, but the trial could not have been completed without MARCC capability and the staff's willingness to accommodate urgent requests for computing time when suitable patients became available, she said.
Video credit: Dave Schmelick and Deirdre Hammer
Mechanical engineer Charles Meneveau, who is working with Hopkins' colleague Dennice Gayme on computer models that can be used to improve wind farm design and operations, echoed those sentiments.
"As a resource, to access MARCC, I feel rich," Meneveau said.
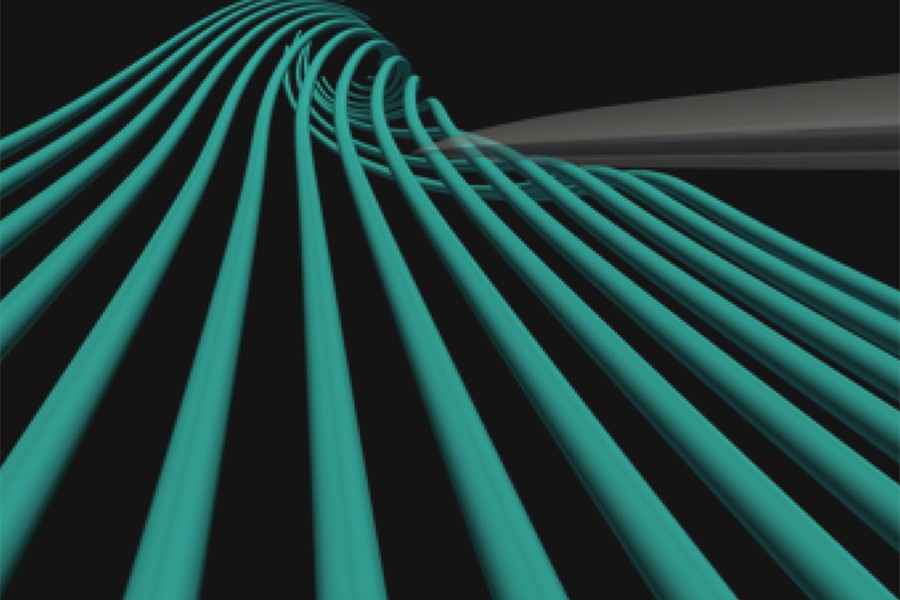
Meneveau's group, before MARCC and since, has produced computer simulations that you can watch on a screen via graphics animation: wind flowing in meandering streams and a scattering of moving streaks over a field of spinning wind turbines. Behind the animations lie months of complex mathematical calculations taking account of prevailing wind and many little twists of air slipping over one turbine blade, moving on and meeting the next.
The model shows the effects of turbulence, meaning wind movement that departs from steady flow. The nuances of turbulence are taken into account in designing turbines and choosing materials best able to stand up to constant buffeting. The work could eventually contribute to wind power development in Maryland and elsewhere.
Rebecca Alford and Jeliazko Jeliazkov, both PhD students in Jeffrey Gray's protein-modeling laboratory, which is part of Whiting's Department of Chemical and Biomolecular Engineering, say MARCC speeds the time-consuming task of producing detailed models of proteins such as antibodies. Any one of the proteins they're simulating—including membrane proteins targeted by more than 60 percent of medications on the market—can take up to 1,000 hours of computing time to complete.
With the advent of MARCC, said Jeliazkov, "we got a million more hours per quarter" of the year.
Posted in Science+Technology
Tagged biomedical engineering, computer science, computer engineering