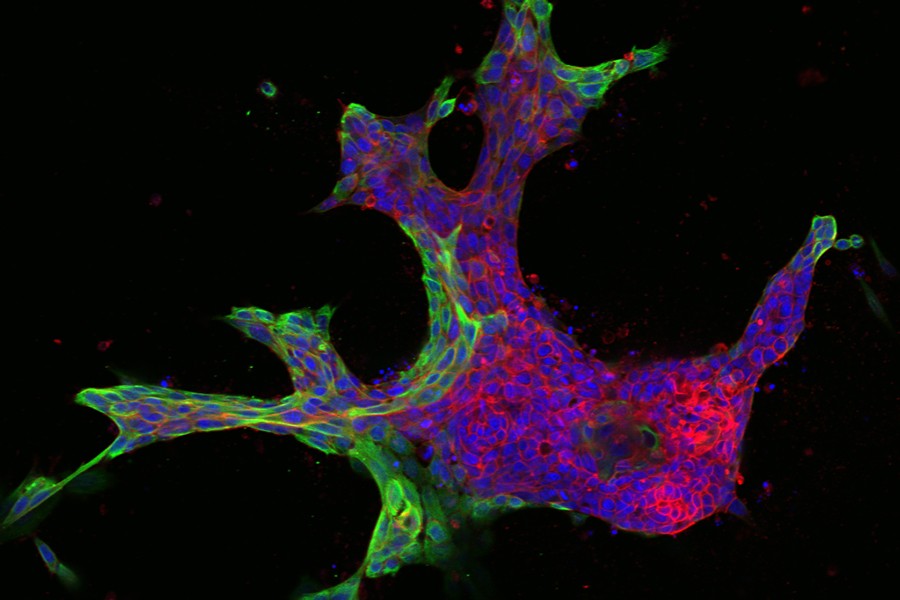
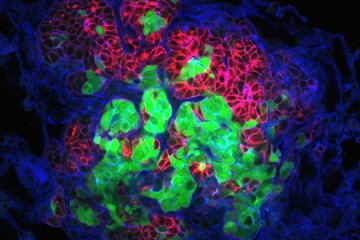
The images were striking—cells clustering into patterns that, to biomedical engineer Joel Bader, looked like starfish and fruits and inkblots.
It was a Johns Hopkins colleague, cell biologist Andrew Ewald, who shared these models with Bader several years ago. The images revealed variations in the patterns of breast tissue grown in Ewald's lab—3-D cultures, or "organoids," derived from the cells of actual cancer patients.

Image caption: Joel Bader (left) and Andrew Ewald
Ewald was looking into the mutations these cells relied upon to spread breast cancer, aiming to pinpoint specific genetic pathways. When Bader came on board, with his background in computational biology, he suggested a numbers-based approach.
"I started thinking that no one had really studied the invasive process quantitatively," Bader says. "The idea was to see which of these different pathways might be significant for people's cancers."
With this groundbreaking strategy—using Bader's computer models to uncover patterns among tens of thousands of Ewald's organoids—the two believe they can help make sense of some of the genetic mysteries behind breast cancer.
Backed now by a $5 million grant from the National Cancer Institute, the pair have joined forces with Johns Hopkins clinicians to dive deeper into this research. The cross-disciplinary effort—called the Johns Hopkins Center for Cancer Target and Development—is now part of a national network of CTD2 research centers.
While traditional efforts to identify breast cancer targets focus exclusively on late-stage tumors, the Hopkins team starts at square one, working with normal breast tissue through its development to a diverse range of tumor stages. They're interested chiefly in the process of metastasis, responsible for the overwhelming majority of deaths from breast cancer.
"Despite its importance, metastasis is not well understood, and the drugs still don't work well enough," Ewald says.
The research relies on live tissue from volunteer breast cancer patients at The Johns Hopkins Hospital, which Ewald's lab uses to generate its 3-D organoids. Each sample can yield thousands of organoids derived from different parts of the tumor, reflecting a complex spectrum of mutations, gene expression, and invasive behavior.
From there, Bader's computer models classify the organoids numerically, matching the varying shapes with their molecular drivers. He starts by ordering the organoids from the simplest to the most complex shapes—beachball-like versus starfish, for example—to rank them from least to most invasive.
Overall, the goal is to zero in on mutations and gene expression patterns that allow breast cancer to spread to distant organs. Once these key molecules are identified, the researchers say, better drugs can be developed to disrupt metastasis.
"We're trying to discover new molecular targets that are important for cancer at different stages of metastasis—for invasion, dissemination, and regrowth," Bader says.
On the clinical end, three Johns Hopkins Medicine faculty members—David Euhus, Ashley Cimino-Mathews, and Edward Gabrielson—help patients donate their tumor samples as part of the surgical process.
The research builds on Ewald's 15 years working with organoids, "trying to make metastasis something we can understand in the lab," he says. For Bader, a theoretical and computational scientist with expertise in genomics, the collaboration has marked his first venture into cancer research.
The two first joined forces in 2012 to identify molecular drivers of cell escape, then continued to study how groups of cancer cells spread through the body. Their early collaborations won grants from the American Cancer Society and the Jane Koskinas Ted Giovanis Foundation to develop the cancer organoids and the mathematical approaches respectively, paving the way for the recent National Cancer Institute grant.
"We're continuing to work toward a better understanding of metastasis," Ewald says. "Once these big questions are unlocked, we'll have a much better shot at improving patients' outcomes with breast cancer."
Posted in Health, Science+Technology
Tagged cancer, breast cancer, andrew ewald, joel bader