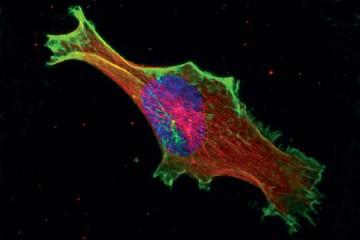
Because of results seen in flat lab dishes, biologists have believed that cancer cells move through the body in a slow, aimless fashion—much like an intoxicated person unable to walk in a straight line.
While this behavior, called a random walk, may hold true for cells traveling across two-dimensional lab containers, researchers at Johns Hopkins have discovered that for cells moving through three-dimensional spaces within the body, the "drunken" model of movement simply doesn't hold true.
This discovery, detailed online this month in Proceedings of the National Academy of Sciences, is important because it should help scientists more accurately predict how cancer will spread through the body. The study's authors have produced a new mathematical formula that they say better reflects the behavior of cells migrating through 3D environments.
The research was supervised by Denis Wirtz, a professor of chemical and biomolecular engineering, materials science, pathology, and oncology. Wirtz said the discovery is part of a current shift toward studying how cells move within three dimensions. In earlier studies, his team showed that cells in 2D and 3D environments behave differently, contrary the previous belief that this behavior is the same, regardless of the environment.
"Cancer cells that break away from a primary tumor will seek out blood vessels and lymph nodes to escape and metastasize to distant organs," Wirtz says. "For a long time, researchers have believed that these cells make their way to these blood vessels through random walks. In this study, we found out that they do not. Instead, we saw that these cells will follow more direct, almost straight-line trajectories. This gives them a more efficient way to reach blood vessels—and a more effective way to spread cancer."
For researchers trying to understand how metastasis occurs, he added, this finding has critical implications.
"This means that the time these cancer cells need to make their way out of connective tissues is much shorter than previous estimates," says Wirtz, who was recently named the university's vice provost for research.
The co-lead authors on the PNAS paper are Pei-Hsun Wu, a postdoctoral fellow, and Anjil Giri, a doctoral student, both in the university's Department of Chemical and Biomolecular Engineering. Wu said the team knew that the persistent random walk model, developed for characterizing cell movements in flat Petri dishes, was also being used in 3D testing, yielding questionable results in the latter.
"It has been used for both kinds of experiments because it's easy and convenient," Wu says. "But it really doesn't fit well when you are working in 3D. Our new math model works better in both 2D and 3D testing."
The researchers discovered that cells in a 3D matrix exhibit different magnitudes of movement in different directions. To address this, the team members enhanced the original formula by identifying the primary and secondary directions in which the cells move, along with the speeds at which the cells travel and their persistence. In cell studies, persistence refers to how far the cells move in a fairly straight line before changing direction.
"Cells that are moving through a 3D environment seem to be more directional than those moving across a flat 2D surface," Giri says. "The unpredictable random walk is not prevalent in a 3D environment."
The team's improved math model for studying cell migration was published with their journal article, and the researchers hope other scientists who are trying to understand and prevent cancer metastasis will quickly adopt it. Although the team members used fibrosarcoma cancer cells in the PNAS study, they said the new model can also be used to help understand the behavior of other cell types, including those that move through the body to help fight infections and to speed the healing of wounds.
Co-authors of the study were Sean X. Sun, an associate professor in the Department of Mechanical Engineering, and Wirtz, who directs the Johns Hopkins Physical-Sciences Oncology Center and is affiliated with the School of Medicine's Sidney Kimmel Comprehensive Cancer Center.
Posted in Health, Science+Technology
Tagged cancer, pathology, biomolecular engineering, denis wirtz