The phrase "sequencing a genome" misleads. It makes the process sound so straightforward: A simple sequence of events leads to a sequenced genome. First, take the DNA that makes up the genome. Spool out the now familiar double helix of the DNA molecule, the orderly spiral ladder of adenine and guanine, cytosine and thymine. Separate the strands, then use a machine to start at one end and record each base—AAGCTAGCTAGC and on and on and on—until you reach the end. Done. No more complicated than reading the digits of pi, except unlike pi the genome is finite, which should make reading it even more straightforward.
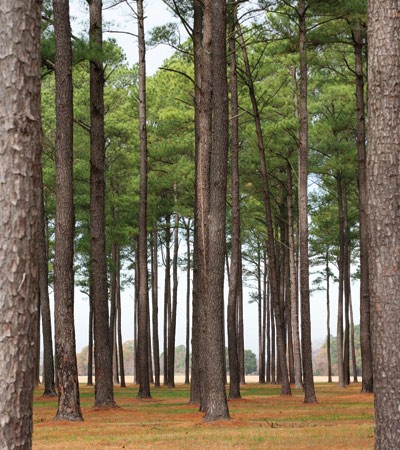

Image credit: Istockphoto.com
Actual sequencing bears little resemblance to that orderly process. The human genome consists of about 3 billion DNA base pairs. Current sequencing technology requires starting with a solution that contains millions of copies of the genome broken into tiny random fragments of roughly 150 base pairs each, the maximum size that the technology can accurately read. This produces hundreds of millions of read bits with no instructions for how to reassemble them into the blueprint for a single molecule.
There are not many people in the world who know how to sift those hundreds of millions of fragmentary DNA reads and assemble a single, accurately sequenced genome. Steven Salzberg, a professor in the McKusick-Nathans Institute of Genetic Medicine at Johns Hopkins, is one of them. He is not a biologist or geneticist but a computer scientist. He and his team create algorithms that sort the innumerable fragments of sequenced DNA and place them in the proper sequence. He got into this line of work while a graduate student in computer science at Harvard (he completed his doctorate in 1989). At the time, the Human Genome Project was just getting under way. "I heard about it and thought, That is going to be the biggest thing in science. I have to see if I can get involved in that," he says. He expanded his studies to include genetics and genomic technology, and identified some problems in DNA sequence analysis to which he thought he could apply his computer expertise. Now he is one of the world's experts in the computational process of genome assembly.
That means he gets calls for interesting projects. After the 2001 anthrax attacks that killed five people in the United States, he was part of a team at the Institute for Genomic Research in Rockville, Maryland, that sequenced the strains of anthrax used in the attacks. He has worked on the mitochondrial DNA of a Columbian mammoth that lived in North America about 11,000 years ago. He is collaborating with Cynthia Sears, an infectious disease specialist at the School of Medicine, to sequence a bacterium associated with colon cancer.
He is also working on the most complicated sequencing yet attempted: the genome of the loblolly pine tree. The loblolly's genome runs to about 22 billion base pairs, roughly seven times longer than the human counterpart. Biologists and agricultural scientists have ample reason for wanting to know what all is in there because the loblolly pine is the most commonly farmed tree in the United States and the second most common species after the red maple. Understand the genome and you have the potential ability to manipulate it to respond to new diseases and environmental changes, and to engineer the crop to be more productive.
Salzberg's team completed the basic assembly in March and is now refining it. They had to piece together about 16 billion separate DNA reads, each one in its proper place. Salzberg characterizes the challenge like this: "Imagine that we have today's newspaper, and suppose we took 100,000 copies and shredded them in such a way that you could read only 100 to 150 letters in a row. You have all these fragments, and now I tell you I want just one reassembled copy of the newspaper." Plus the data obtained from the sequencer is not free of noise. The many copies of DNA used in the process can contain slight variations, and the sequencing process introduces errors at a rate of 0.5 percent. In the course of 16 billion reads, those errors add up.
To further complicate matters, genomes are filled with repetitive sequences. For example, the human genome has sequences 300 bases long that occur, repeated almost exactly, more than a million times. That means if you take any one fragment, there may be a million places where it could go. "It's like the blue sky section of a jigsaw puzzle," Salzberg says. "If all the pieces are the same color, you don't know where they go. It's actually much worse than that. Imagine if all the pieces were the same shape, too."
Nobody knows why the loblolly genome is so immense. "Pine trees are not very smart, yet their genomes are seven times bigger than ours. They don't even have a brain! How come they have a genome that is seven times bigger?" Salzberg asks. Even the amoeba has a larger genome than humans. Why? "It's good cocktail party conversation," Salzberg says. "We don't really know the answer."
Posted in Science+Technology